Principal Investigator: Dr. Tom Warkentin
Co-investigators: Dr. Krishna Kishore Gali, Nimllash T. Sivachandra Kumar, Dr. Sabine Banniza
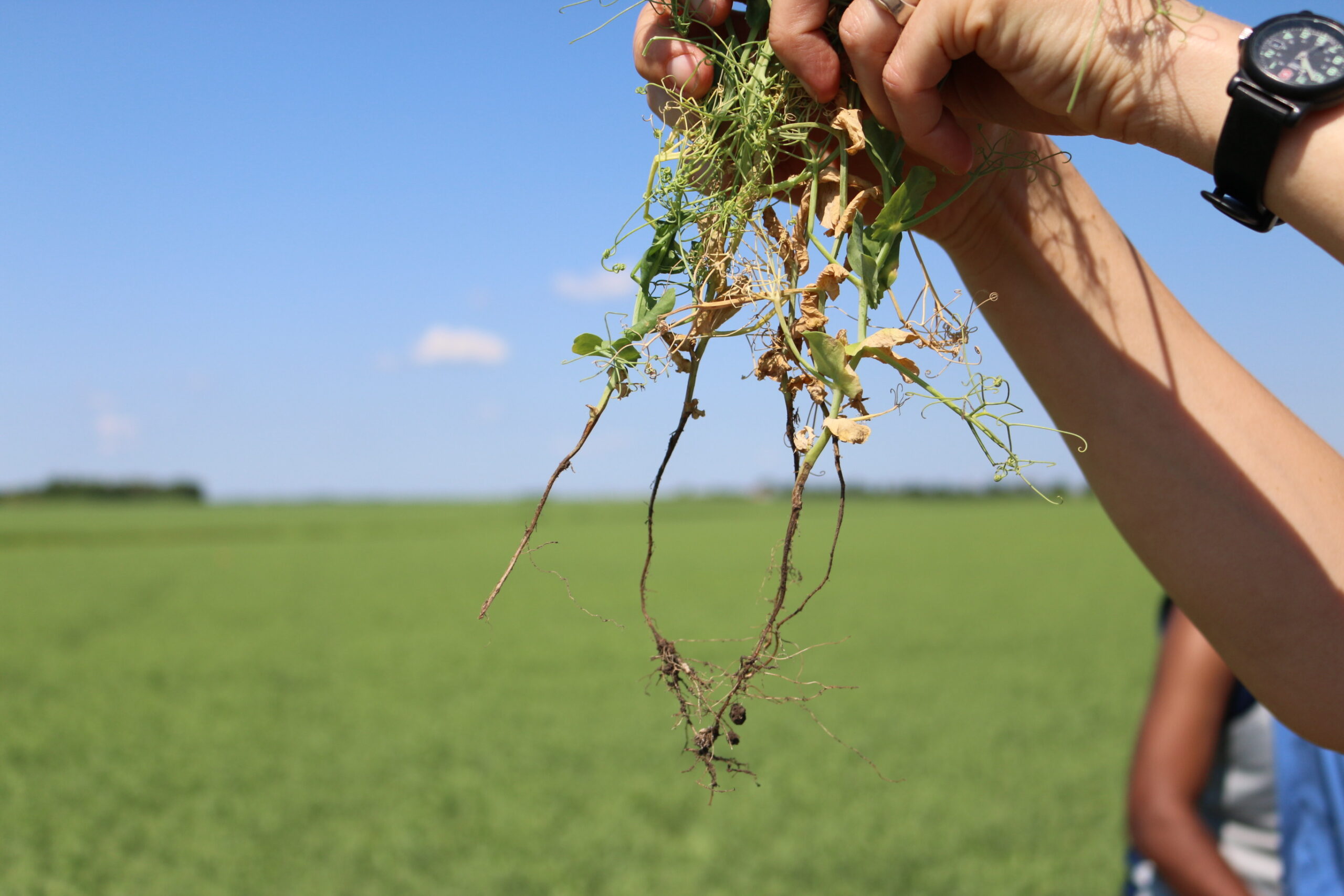
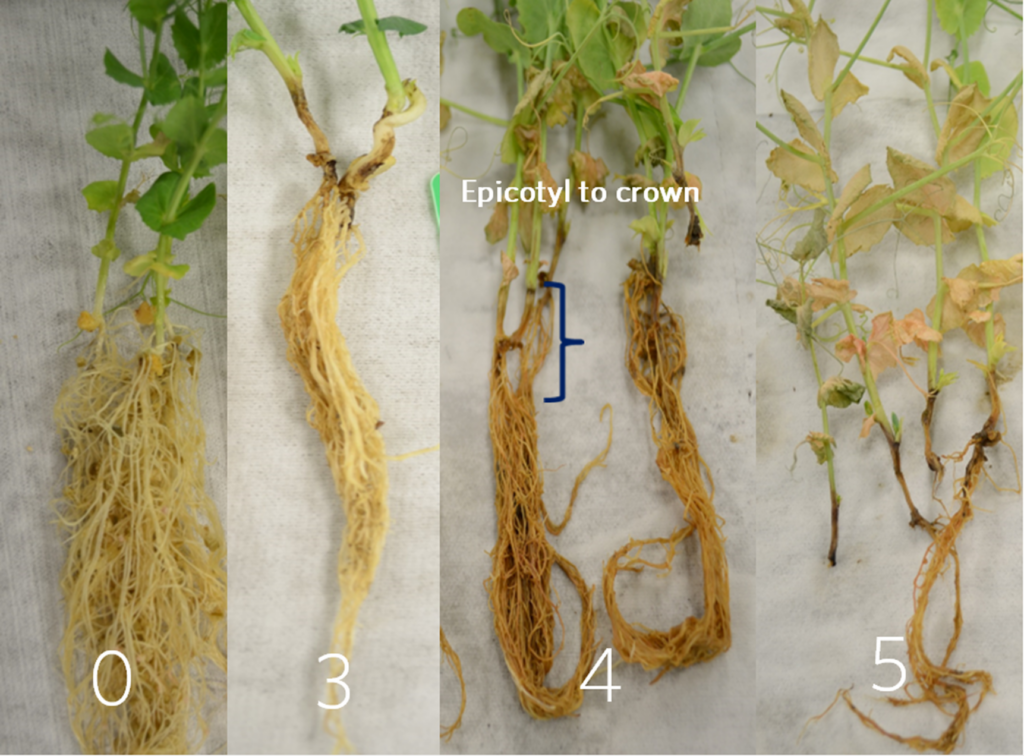
The oomycete pathogen Aphanomyces euteiches is the causal agent of Aphanomyces root rot (ARR), a destructive soilborne disease of pea (Pisum sativum). ARR leads to root browning, cortical decay, and reduced nodulation (Figure 1), ultimately causing significant yield losses in pea-growing regions worldwide due to the pathogen’s long-lived oospores and persistence in agricultural soils.

In response to the persistent threat of ARR, researchers in the United States (U.S.) and France have invested substantial resources in breeding programs to develop ARR-resistant pea cultivars. These efforts integrate field screening in infested soils, quantitative trait locus (QTL) mapping, and marker-assisted selection to accelerate the deployment of resistant germplasm and improve long-term disease management strategies. However, a lack of adequate levels of disease resistance in Canadian cultivars is concerning for pea production in Western Canada.
Following fundamental research by the United States Department of Agriculture (USDA) and French researchers, pea lines with improved resistance have been identified, and two QTLs associated with resistance have been identified: Ae-Ps7.6 and Ae-Ps4.5. The Crop Development Centre (CDC) pea breeding and pathology programs have used a marker-assisted backcross breeding approach to incorporate partial resistance identified in the U.S. and France into adapted CDC varieties. Four elite lines, pyramided with QTLs Ae-Ps7.6 and Ae-Ps4.5, were tested for disease resistance in controlled-environment disease resistance trials and for agronomic performance in yield trials, and they were entered in variety registration trials beginning in 2023.
However, recent research from the Institut national de la recherche agronomique (INRA) and the University of Rennes in France has shown that high selection pressure can rapidly erode currently available resistance; that is, resistance based on limited genetic sources is reduced under sustained selection. Growing pea lines with the identified resistance alleles consecutively in the same field for four years increased the proportion of more virulent pathogenic isolates by 18%. Furthermore, forty-three A. euteiches isolates from different French pea-growing regions, when tested against the resistance allele at QTL Ae-Ps7.6, exhibited high, moderate, and low aggressiveness, suggesting a future risk of erosion of the QTL effect.
There is also evidence that Canadian isolates of ARR can be linked to a single source of resistance, and this narrow resistance base has raised concerns about long-term resistance and durability under prairie conditions. Therefore, there is an ongoing need to identify new and diverse sources of resistance in pea, as well as to stack multiple resistant genes into the same pea variety. Recently, two new QTLs were identified by researchers at the CDC at the University of Saskatchewan (USask) as important for imparting ARR resistance against Saskatchewan isolates of A. euteiche, Ae-Ps1.2, and Ae-Ps4.1 s (Gali et al., 2025). Furthermore, the CDC at the USask, developed a genome wide resistance association panel (termed GWAS-2) which is a collection of 233 diverse pea core germplasm and breeding lines for potential identification of new sources of ARR resistance and additional QTLs that might be effective in breeding for ARR resistance in Western Canada, as well as 39 P. fulvum (wild pea) lines for ARR resistance.
Thus, this current project aimed to test the potentially new and diverse sources of pea ARR resistance screened under the GWAS-2 pea panel using four objectives:
- Identify resistant strains of pea by phenotyping the GWAS-2 pea panel for ARR resistance.
- Of the identified resistant germlines, conduct association mapping of the genome to identify genetic markers associated with ARR resistance.
- Computationally compare the identified genetic markers with known QTLs of ARR resistance and identify new candidate genes.
- Field evaluate the selected GWAS-2 germlines to resistance to ARR.
Objective 1: Phenotyping the GWAS-2 pea panel for ARR resistance
The GWAS-2 panel was evaluated for resistance to A. euteiche under controlled growth-chamber conditions using replicated randomized complete-block designs. Seedlings were inoculated with a standardized zoospore suspension, and disease severity was scored using a quantitative rating scale from one to 7. Symptoms were converted to a numerical value based on the following root discoloration symptoms: (1) indicating no root discoloration, (2) 1–20% discoloration, (3) 21–40% discoloration, (4) 41–60% discoloration, (5) 61–80% discoloration, (6) 81–100% discoloration, and (7) dead plant. The disease ratings were then converted to disease severity using 0% for a score of 1, 100% for a score of seven and the mid-point values of percent root discoloration for scores of two to six (2 = 10%; 3 =30%; 4 = 50%; 5 = 70%; 6 = 90%). Statistical analyses were conducted using generalized mixed models to ensure appropriate model fit. Additionally, wild P. fulvum lines were screened using a similar protocol to identify potential novel sources of resistance.
ARR severity among the GWAS-2 pea accessions ranged from 36.1% to 84.9%, indicating substantial variation in resistance (Figure 2). The most resistant line identified was AAC Ardill with a disease severity rating of 36.1%, followed by PI 639976 (disease severity 37.8%) and Austrian Winter type pea Melrose (disease severity 38%), all performing substantially better than the susceptible check CDC Meadow (disease severity 81.8%) and comparable to known moderately resistant checks PI 660736 (disease severity 32.6%) and PI 660729 (disease severity 43.7%). In total, ten accessions were identified as new sources of moderate resistance, expanding the available genetic diversity for breeding. Additionally, two wild P. fulvum accessions (PI 260064 and PI 560065) exhibited moderate resistance, with disease severity ratings of 45% and 48.6%, respectively. These findings significantly broaden the genetic base for ARR resistance and provide valuable material for developing pea varieties with more durable disease resistance.
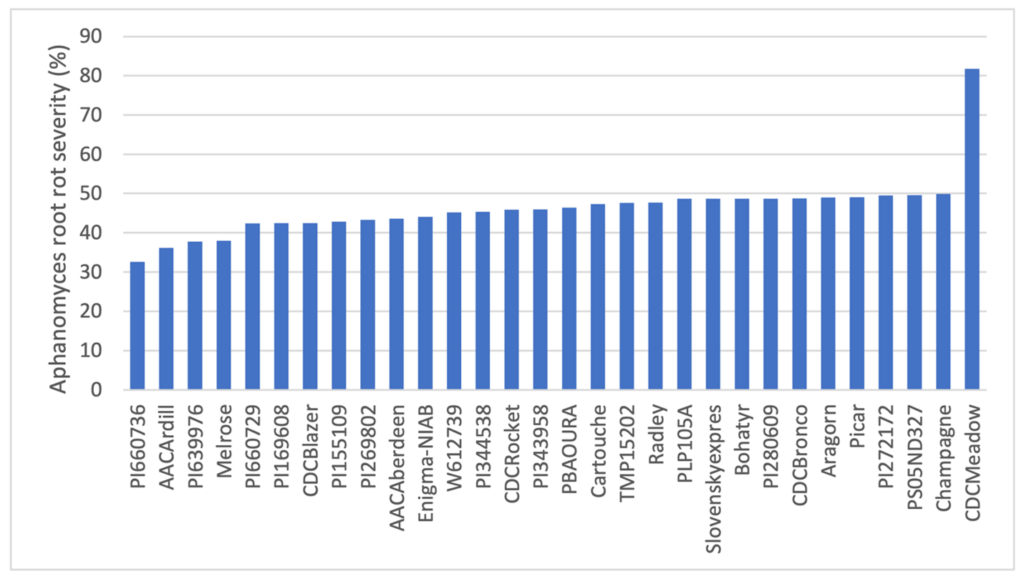
Source: Tom Warkentin, CDC, Department of Plant Sciences, USask.
Objective 2: Association mapping to identify SNP markers linked to ARR resistance
The GWAS-2 panel was genotyped, and high-quality filtered markers were used for population structure and genome-wide association analyses. Multiple GWAS models were compared to identify robust marker–trait associations, resulting in genetic markers significantly associated with ARR resistance.
Genetic analysis grouped the GWAS-2 pea accessions into 14 population clusters. Genome-wide association mapping identified several DNA markers that were significantly associated with ARR resistance across multiple chromosomes. The most significant marker, Chr7LG7_429174184, explained 57.6% of the variation in disease resistance, followed by Chr4LG4_206450021. Additional significant markers were also identified on chromosomes 1, 2, 3, and 6. This will allow researchers to pinpoint genetic signposts in peas that will help determine if the plant can better withstand ARR and provide multiple targets for marker-assisted breeding.
Objective 3: Computational comparison of identified gene markers with known QTLs and candidate gene identification
Of the gene markers identified through the GWAS2 panel, the two best markers associated with ARR resistance were compared with previously reported ARR resistance QTLs using the pea reference genome sequence to identify co-localized genes. Genomic regions surrounding the most significant markers were also examined, ±500 kb in either direction, to identify further candidate genes associated with disease resistance.
Comparison with previously reported resistance regions showed that the two most significant gene markers, Chr7LG7_429174184 and Chr4LG4_206450021, are located at substantial distances from known major QTLs, indicating they represent novel resistance loci for ARR. Within 500 kb of these markers, several candidate genes were identified. Notably, Psat7g213880, encoding a Cupin-like domain protein linked to plant defence, was located near Chr7LG7_429174184, and Psat4g110720, an NmrA-like family gene associated with stress and defence responses, was located very close to Chr4LG4_206450021. These findings suggest that the genomic regions surrounding these markers likely contain functional resistance genes and provide strong targets for future validation and marker-assisted breeding efforts.
Objective 4: Field evaluation of selected accessions for resistance to the root rot complex
In summer 2024, 233 GWAS-2 pea accessions were evaluated under field conditions in a root rot nursery containing multiple soilborne pathogens. At approximately five weeks post-planting, the plants were evaluated for disease symptoms to confirm that resistance observed in controlled experiments also performs well under real-world growing conditions. Eight to ten plants from each row were uprooted, and the roots were washed and scored for root rot symptoms using a similar 1–7 rating scale as objective 1, with (1) indicating no lesions; (2) 1–2 mm brown lesions at seed attachment; (3) coalescing of localized root/seed attachment lesions approximately 180◦ around the stem, with lesions from 5–10mm; (4) lesions extending and completely encircling the tap root; (5) increasingly discolored and extending to epicotyl; (6) epicotyl lesions extending to green stem and scale leaf; (7) tap root completely brown/black.
Nearly all lines showed severe disease symptoms, with ratings between five and seven on a 1–7 scale. Only one germline, W6 44573, demonstrated moderate resistance (rating 3.8). In controlled environment tests, ARR severity for W6 44573 was rated at 62%, compared to 82% for susceptible check CDC Meadow. This line has also shown moderate resistance in controlled Aphanomyces tests and to Fusarium avenaceum in parallel studies. Despite this observation, there was little overlap between lines that performed well in controlled Aphanomyces assays from objective two and those that showed resistance in the field nursery (Figure 3). These results were likely due to the field site containing multiple A. euteiches pathogens, of which the plants identified as resistant in the controlled setting may not also be resistant to.
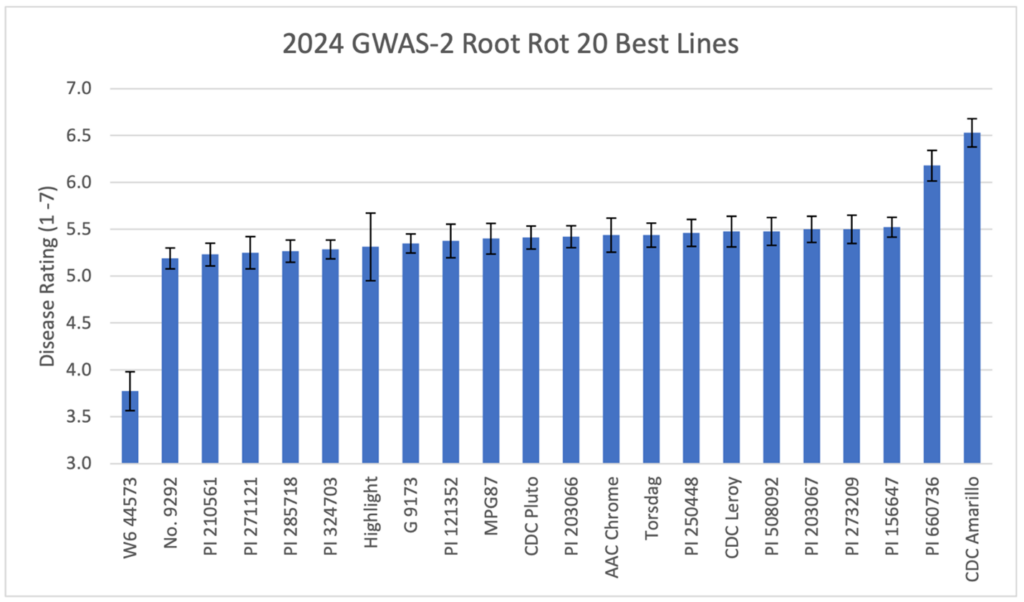
Source: Tom Warkentin, CDC, Department of Plant Sciences, USask.
Conclusions and Implications to Growers
Overall, screening the GWAS-2 pea panel identified ten new sources of moderate resistance to ARR, including AAC Ardill, Melrose, and CDC Blazer, with resistance levels comparable to those of known moderately resistant varieties. Genetic analysis also identified two new DNA markers, Chr7LG7_429174184 and Chr4LG4_206450021, linked to root rot resistance and amenable to marker-assisted breeding to accelerate the development of resistant pea varieties. Importantly, these markers were located at substantial distances from known major QTLs, indicating novel sources of resistance through the candidate genes identified near the markers. Additionally, two wild pea accessions (P. fulvum PI 260064 and PI 560065) were identified with moderate resistance, offering valuable new germplasm for breeding programs.
While field evaluation revealed only one germline exhibiting moderate resistance to root rot under multiple pathogen pressures, the findings from this study have identified novel resistance markers that can be used in future breeding programs. These results directly support new research directions. Future research is already underway, and to combine these resistance traits with other desirable traits, an eight-parent Multiparent Advanced Generation Inter-Cross (MAGIC) population was developed, including the new resistant lines and elite cultivars, with ongoing characterization to support long-term improvement of disease-resistant peas, supported by Genome Canada and Genome Alberta.
For growers, these results represent an important step toward more reliable and durable root rot resistance in future pea cultivars. The identification of novel resistance loci will help breeders develop varieties with broader and more stable protection against Aphanomyces and the root rot complex, reducing yield losses and improving stand establishment in infested fields. Over time, the deployment of pyramided resistance is expected to enhance the sustainability of pea production in Western Canada and reduce the risk of resistance breakdown under continuous cropping systems.



